|
10/18/2023 0 Comments Building blocks of dna and rna 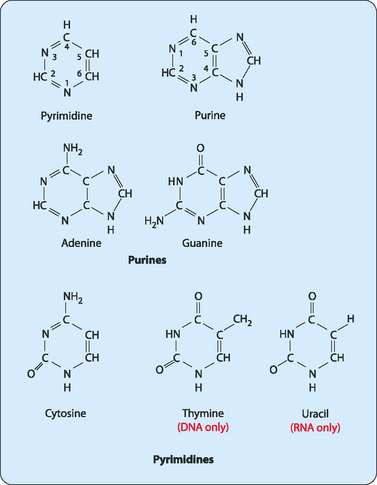 
High-resolution crystal structures were determined for three different hachimoji duplexes assembled from three self-complementary 16-mer sequences: 5’-CTTATPBTASZATAAG, 5’-CTTAPCBTASGZTAAG, and 5’-CTTATPPSBZZATAAG. These errors were said to be similar to those observed with nearest-neighbor parameters for standard DNA:DNA duplexes, which was interpreted as meaning that “GACTZPSB hachimoji DNA reproduces, in expanded form, the molecular recognition behavior of standard 4-letter DNA. Plots of experimental versus predicted free-energy change (ΔG° 37) ( A) and experimental versus predicted melting temperature (T m) ( B) shown here indicate that, on average, T mis predicted to within 2.1☌ for the 94 GACTZPSB hachimoji duplexes, and ΔG° 37 is predicted to within 0.39 kcal/mol. These experimental T mvalues were then compared to predicted melting temperatures derived from state-of-the art thermodynamic parameterization of nearest-neighbor base-pair dimers, as described by Hoshika et al. Duplexes of these GACTZPSB-containing hachimoji 8-mers were then used to measure melting temperature (T m) values under a set of standard conditions. The nucleotides Z, P, S and B along with A, G, C and T were each incorporated into 94 different 8-mer sequences of hachimoji DNA oligonucleotides by use of otherwise conventional phosphoramidite chemistry for solid-phase chain-assembly. Copyright © 2019, American Association for the Advancement of Science, with permission.Įxpansion from six letters to eight letters was investigated using two additional nucleotides, S and B, shown here. These Z and P nucleotides were shown to undergo enzymatic copying, PCR amplification, and successive transcriptions, first into six-letter RNA, and then back into six-letter DNA. The H-bonding in these structures is between oppositely positioned donor (red) and acceptor (blue) atoms. coined this term by combining the Japanese words for eight (“hachi”) and letters (“moji”).Īt the outset, I should point out that there is a YouTube video lecture by Benner that is well worth watching to fully appreciate the rationale behind investigating AEGIS, and the experimental approaches explored.īenner’s previously published work ( Zhang et al.) on the evolution of a functional six nucleotide genetic code included the two new nucleotides, Z and P, which are shown below for DNA dR is replaced with R in RNA. This recently expanded system is appropriately named “hachimoji” DNA and RNA: Hoshika et al. This blog will highlight Benner’s 2019 Sciencepublication ( Hoshika et al.) on AEGIS, which reports further expansion to eight building-block letters. Benner on expanding the genetic code from four to six building-block letters is reviewed therein. Res.by Richards and Georgiadis, titled Toward an Expanded Genome: Structural and Computational Characterization of an Artificially Expanded Genetic Information System(AEGIS). Readers interested in an overview these subjects can consult a 2017 review in Acc. 
Exploring the expansion of such H-bonding (shown here) to include synthetic analogs of these four natural nucleobases has been of interest for theoretical and evolutionary reasons, and could have utility for many hybridization-based applications, as well as for storage of information. This information is in “Watson-Crick” DNA for replication, transcribed into RNA, and finally decoded into protein. In 2019, Coding Uses Eight (“Hachi”) Letters (“Moji”)Īccording to the principles attributed to the early writings of Francis Crick in 1958, The Central Dogma of Molecular Biologystates that H-bonding between A/T and C/G base pairs underlies the storage of genetic information.Benner Led Expansion from Four to Six Letter-Coding in 2015 Researchers Seek to Expand A, G, C, and T Genetic Coding to Additional Nucleobase “Letters”.When they are bound together by mRNA, they form a polysome. Each molecule of mRNA is long enough for 6 to 10 ribosomes to attach to it and read it simultaneously. 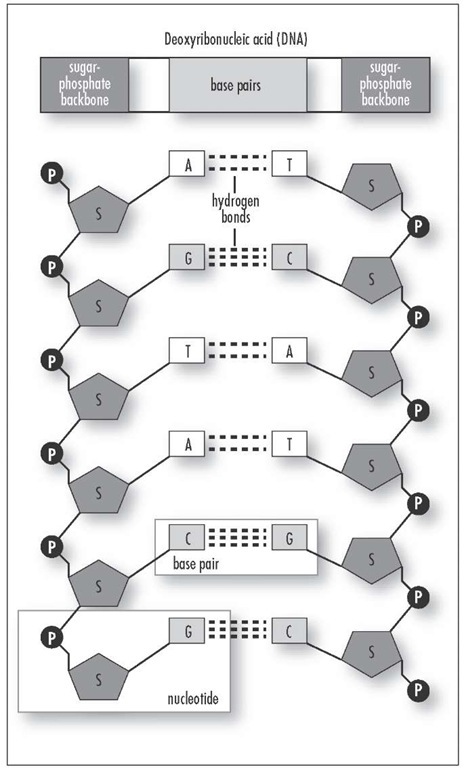
While most plants synthesize few proteins and have few ribosomes, some others produce more like the protein-rich seeds of legumes. The ribosome in plants is usually made of three molecules of rRNA and about 50 types of proteins that associate and form into subunits. In plants, the ribosome can be within the Mitochondria and Plastids. On arrival, the SRP is released and protein synthesis starts up again." "The ribosome halts protein production while the SRP brings the ribosome and its partly-built protein to where it’s needed in the cell. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
